Eliminate the mystery surrounding the human nervous system!
Students often dread beginning their study of the nervous system. The nervous system seems so complex and mysterious.
But you should not let the nervous system worry you. The design of its basic circuits for interacting with the body’s ever-changing environment is straightforward.
First, this system has a multitude of specialized sensing units. These sensing units deliver their observations to several central processing units that lie deep in the brain.
Next, the central processing units engage in crosstalk to compare incoming information from multiple sensors. They sort the information and forward it to the response areas distributed across the surface of the brain.
The surface areas of the brain communicate with each other and implement a coordinated response. The response alters input to the sensory units, which in turn relate the outcome of the response back to the neural processing units. It is a continuous cycle.
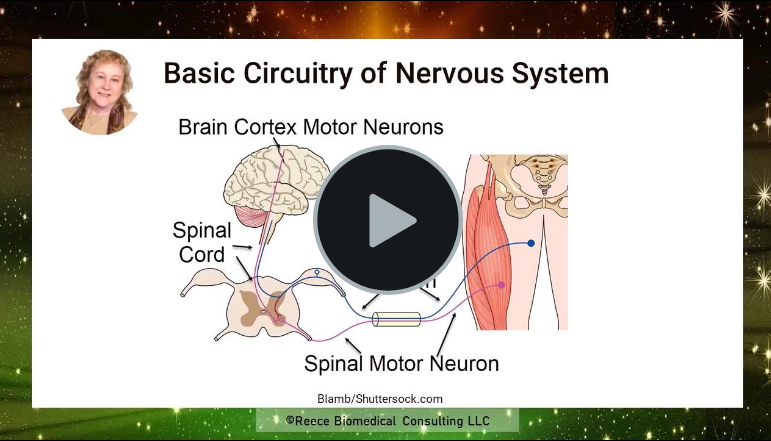
To fill in some of the details of this process click on the play button for this 12-minute video.
Fight-or-Flight response of the human nervous system
The final image in the video displayed below is a good example of the fight-or-flight response to danger. I used this as a summary image in the video, because it provides me an opportunity to synthesize the concepts of sensory input to the brain and motor output by the brain to skeletal and smooth muscles under extreme circumstances.
An avoidance of danger depends upon recognizing the situation as dangerous. If the visual and/or auditory sensory information aligns with a memory of trouble, a deep brain nucleus named the amygdala takes over the physiologic response. The amygdala begins the response before the information gets to the frontal decision-making cortical neurons.
The amygdala works with another group of deep brain nuclei named the hypothalamus to signal the adrenal glands to release their hormones, cortisol and epinephrine. These hormones enhance skeletal muscle speed of contraction and endurance.
Signaling by brain stem nuclei activates the sympathetic nervous system increasing heart and respiratory rate. This enhances delivery of needed nutrients and oxygen to the contracting muscles.
Sympathetic nervous system activity also shunts blood flow to skeletal muscle by contracting the smooth muscle of the arteries elsewhere in the body.
Further Reading
Neurons and Brain’s Other 90% of Cells
Do you have questions?
Please put your questions in the comment box or send them to me by email at DrReece@MedicalScienceNavigator.com. I read and reply to all comments and email.
If you find this article helpful share it with your fellow students or send it to your favorite social media site by clicking on one of the buttons below.
Margaret Thompson Reece PhD, physiologist, former Senior Scientist and Laboratory Director at academic medical centers in California, New York and Massachusetts is now Manager at Reece Biomedical Consulting LLC. She taught physiology for over 30 years to undergraduate and graduate students, at two- and four-year colleges, in the classroom and in the research laboratory. Her books “Physiology: Custom-Designed Chemistry”, “Inside the Closed World of the Brain”, and her online course “30-Day Challenge: Craft Your Plan for Learning Physiology”, and “Busy Student’s Anatomy & Physiology Study Journal” are created for those planning a career in healthcare. More about her books is available at https://www.amazon.com/author/margaretreece. You may contact Dr. Reece at DrReece@MedicalScienceNavigator.com, or on LinkedIn.
She taught physiology for over 30 years to undergraduate and graduate students, at two- and four-year colleges, in the classroom and in the research laboratory. Her books “Physiology: Custom-Designed Chemistry”, “Inside the Closed World of the Brain”, and her online course “30-Day Challenge: Craft Your Plan for Learning Physiology”, and “Busy Student’s Anatomy & Physiology Study Journal” are created for those planning a career in healthcare. More about her books is available at https://www.amazon.com/author/margaretreece. You may contact Dr. Reece at DrReece@MedicalScienceNavigator.com, or on LinkedIn.
Dr. Reece offers a free 30 minute “how-to-get-started” phone conference to students struggling with human anatomy and physiology. Schedule an appointment by email at DrReece@MedicalScienceNavigator.com.